
RT-q PCR REAGENTS
www.clontech.com
Clontech Laboratories, Inc. • A Takara Bio Company
United States/Canada: +1.800.662.2566 • Asia Pacific: +1.650.919.7300 • Europe: +33.(0)1.3904.6880 • Japan: +81.(0)77.543.7247
PrimeScript
™
Reverse
Transcriptase
R
PrimeScript Reverse Transcriptase is a recombinant, RNase H Minus, modified MMLV
(Moloney Murine Leukemia Virus) enzyme developed by Takara Bio. PrimeScript RT is robust
and versatile, and is well-suited for all applications that require full-length first strand cDNA.
PrimeScript RT offers:
• Strong strand displacement and extension capability: Synthesizes long, full-length cDNA
molecules (up to 12 kb)
• High specificity: Capable of synthesizing cDNA at 42°C, which allows high specificity, high
yield, and reduced risk of RNA degradation
• Outstanding accuracy: Low error rate compared to other commercially available reverse
transcriptases
• Fast reactions: Reverse transcription reactions can be completed in as little as 15 minutes
Why does PrimeScript RT provide out-
standing performance?
RNA secondary structure can interfere with
cDNA synthesis. Conventional RTs can anneal
nonspecifically to higher order structures of
RNA. Nonspecific products derived from such
mispriming can impair overall RT efficiency
and reduce the yield of full-length cDNA.
To minimize these effects, reactions with
conventional RTs are frequently performed at
higher temperatures so that RNA secondary
structures are partially or completely denatured.
However, because high temperatures also
increase the risk of RNA degradation, these
reaction conditions are not ideal. In contrast,
PrimeScript RT can be used at 42°C.
PrimeScript RT reduces mispriming events by
avoiding nonspecific annealing and preventing
primer-dimer formation. Furthermore, the
enzyme has strong strand displacement activity.
This tech note introduces the key features of
PrimeScript RT and provides experimental
examples for PrimeScript RT and kits.
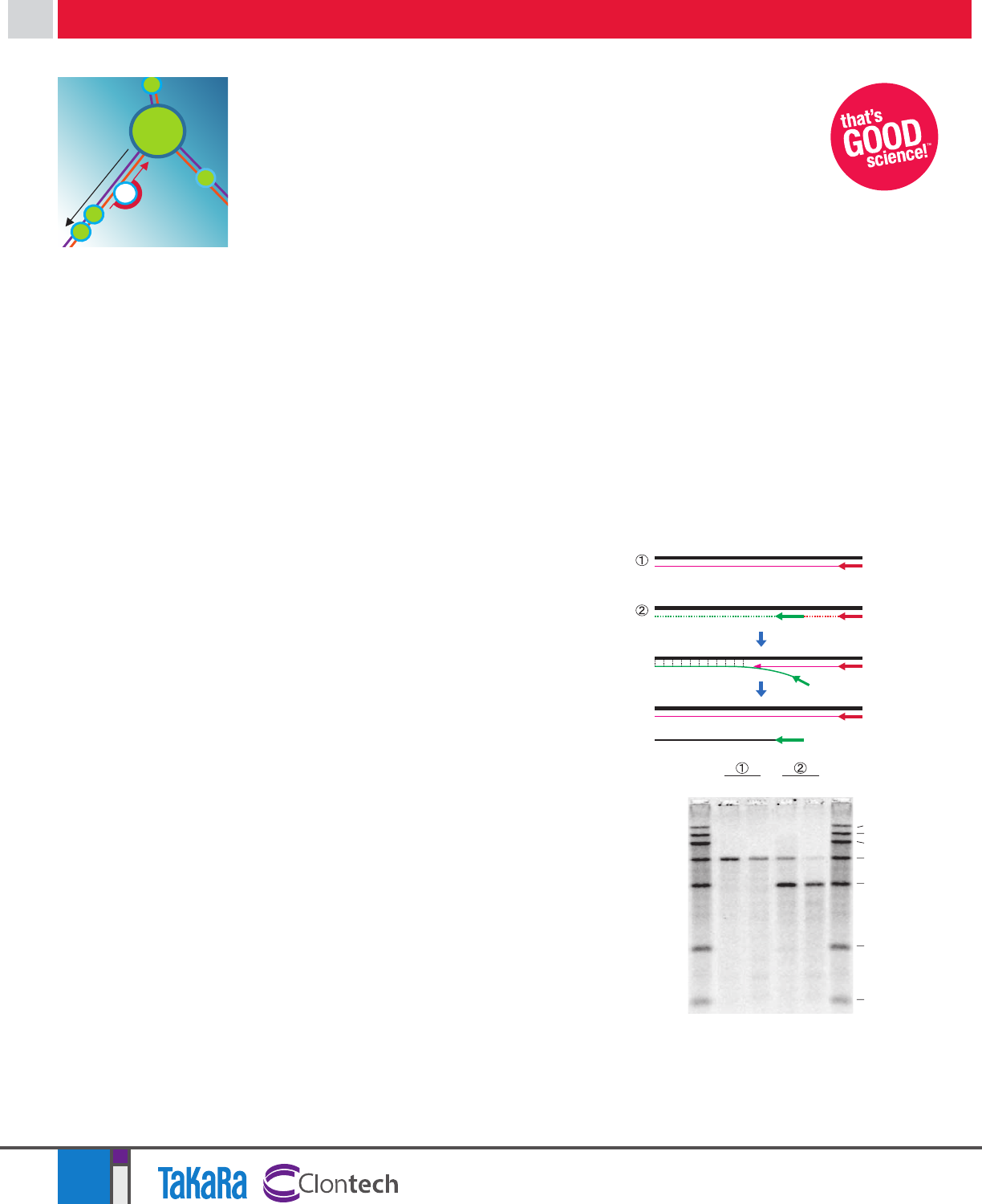
Strong strand displacement activity
of PrimeScript RT results in excellent
extension of long targets
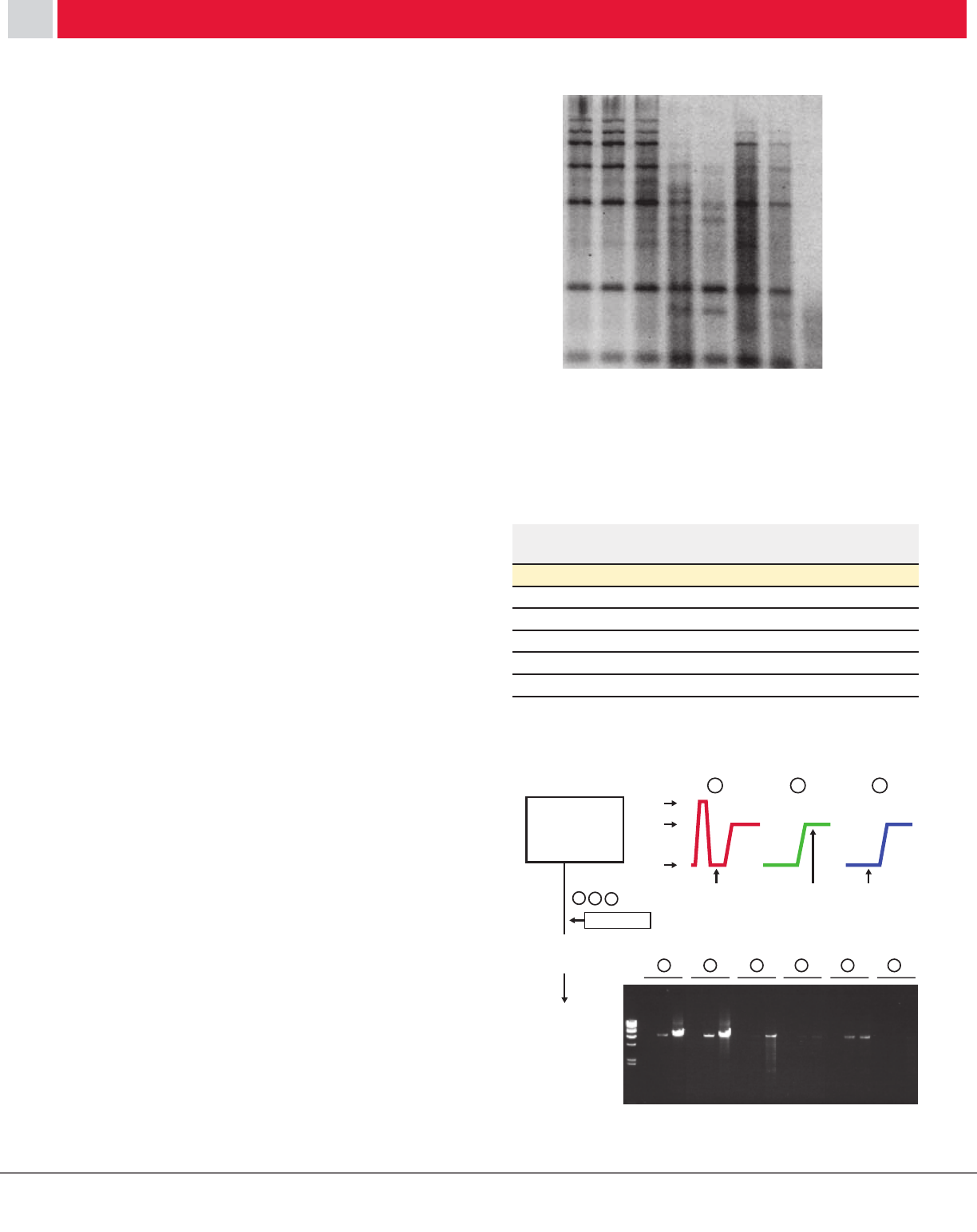
cDNA was prepared by primer extension using
either PrimeScript RT or an RT from Company L
at 50°C. Products originating from an oligo-dT
primer and an internal, specific primer were
analyzed on an alkaline denaturing gel (Figure 1).
Extending the Boundaries
of Reverse Transcription
For Research Use Only. Not for use in diagnostic or therapeutic procedures. Not for resale. SYBR is a registered trademark of Molecular Probes, Inc. TaqMan
is a registered trademark of Roche Molecular Systems, Inc. Takara, the Takara logo, and TaKaRa Ex Taq are trademarks of TAKARA HOLDINGS, Kyoto, Japan.
CellAmp, Premix Ex Taq, PrimeScript, PrimeSTAR, and Thermal Cycler Dice are trademarks of Takara Bio, Inc. Clontech, the Clontech logo, and that’s GOOD
science! are trademarks of Clontech Laboratories, Inc. All other marks are the property of their respective owners. Certain trademarks may not be registered
in all jurisdictions. Clontech is a Takara Bio Company. ©2015 Clontech Laboratories, Inc.
0415 IN 633429
6.4 kb Transcript
6.4 kb
6.4 kb Transcript
4.4 kb
Specific Primer
6.4 kb
4.4 kb
Oligo dT Primer
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
Oligo dT Primer
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
Oligo dT Primer
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
Oligo dT Primer
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
T
A
1 2 1 2
1 : PrimeScript RT
2 : Company L MMLV RT III
kb
12
10
8.4
6.4
4.4
2.0
1.0
Figure 1: Comparison of strand displacement
and elongation activity.
RT-q PCR REAGENTS
2
PrimeScript RT synthesizes cDNA at 42°C with
high specificity and yield
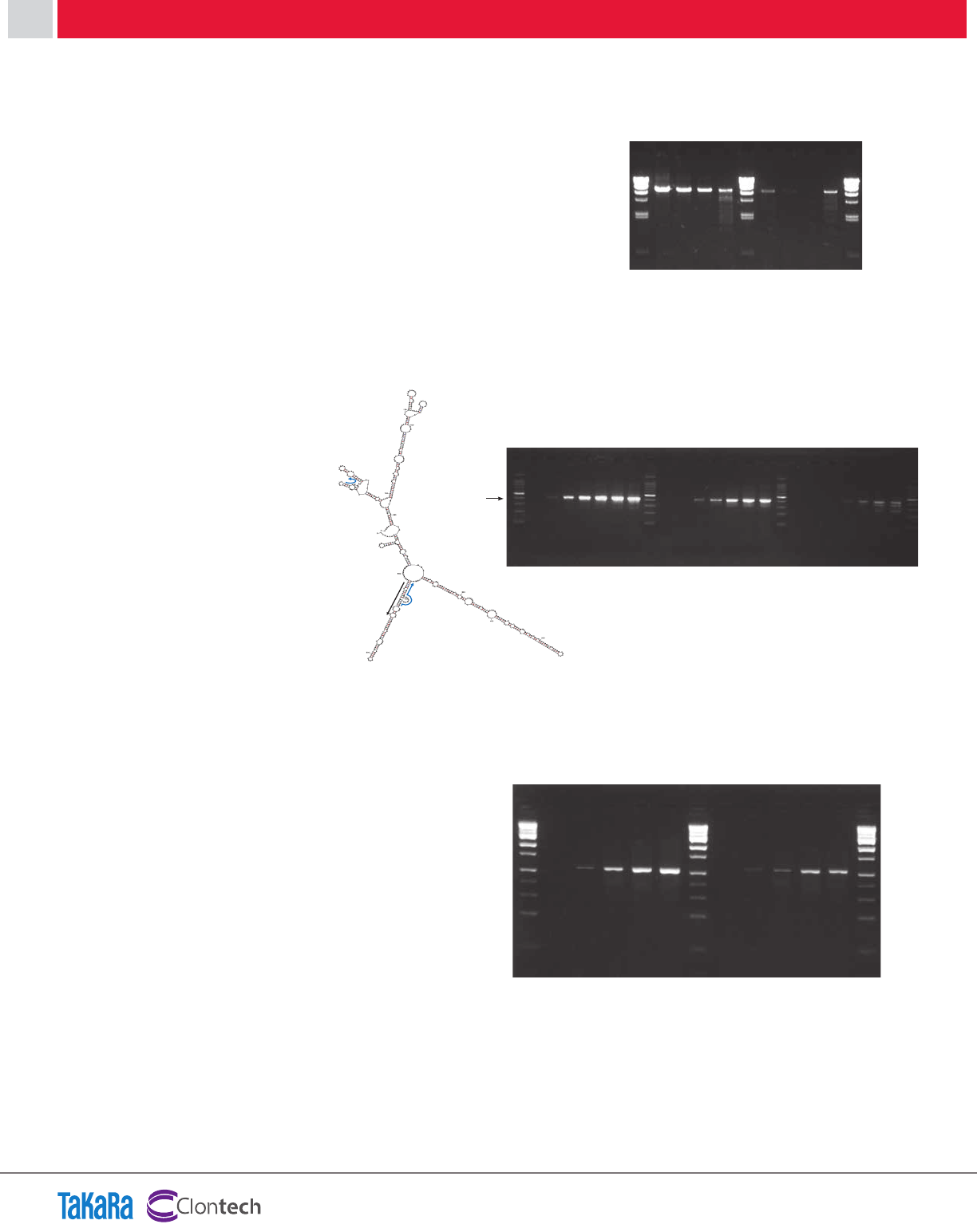
Extension products obtained with PrimeScript RT were
compared with products generated by six other commer-
cially available RTs. An RNA ladder (containing 1, 2, 4.4, 6.4,
8.4, 10, and 12 kb fragments) was used as the template for
first strand cDNA synthesis. Each reaction was assembled
and performed according to the manufacturer’s recommen-
dations. Equivalent amounts of cDNA were loaded onto
an alkaline denaturing gel. After electrophoresis, the gel
was stained with SYBR
®
Green II, and the products were
detected by fluorescence imaging (Figure 2).
The yield of full-length cDNAs was far higher for Prime-
Script RT than any of the other enzymes. In addition,
although the reaction temperature for PrimeScript RT was
lower (42°C), the resulting cDNA had significantly less back-
ground arising from non-specific priming events.
PrimeScript RT is highly accurate
Several different RTs were used to synthesize first-strand
cDNA from human placenta total RNA (Clontech
®
Cat.
#636527) using oligo-dT primers. Reactions were per-
formed following each manufacturer’s recommended
protocol. After cDNA synthesis, PCR amplification of the
TF gene was performed with high-fidelity PrimeSTAR
®
HS
DNA Polymerase (Clontech Cat. # R010A). The 500 bp ampli-
fied fragments were then cloned into vectors, and multiple
clones were chosen for DNA sequence analysis. The error
rate was defined as the number of errors per total number
of bases sequenced (~200,000 bases). A PCR fragment
directly amplified from human genomic DNA was used as
a control. PrimeScript RT resulted in only 7 errors out of
200,000 bases (an error rate of just 0.0035%), the highest
accuracy of all RTs analyzed (Figure 3).
PrimeScript RT facilitates easy workflow
During synthesis of long RNA products by reverse
transcription, conventional enzymes require that the RNA
template be denatured first, followed by addition of RT. This
is intended to avoid the suppression of cDNA synthesis
that can occur when RT anneals to double-stranded RNA
secondary structures. In contrast, PrimeScript RT does not
require template denaturation, allowing greater flexibility in
workflow.
The yield of an 8 kb dystrophin cDNA product produced
by PrimeScript RT and Company L’s RT was examined by
RT-PCR. Three different protocols were compared as shown
in Figure 4: 1) denaturation of RNA template followed
by addition of RT on ice; 2) pre-incubation at the RT
reaction temperature (42°C for PrimeScript RT or 50°C for
Company L’s RT) followed by addition of RT at the reaction
temperature, but without prior template denaturation; or
3) addition of RT to template on ice, then shifting to the
reaction temperature. While PrimeScript RT produced cDNA
with all protocols, Company L’s RT generally showed poor
yield and absence of product when RT was added without
prior template denaturation (protocol 3).
PCR
Protocol
Oligo dt primer
dNTP
Total RNA
(0.1; 1; 10 ng)
RT (15-30 min)
RT inactivation
Target:
Dystophin 8 kb
Denature RNA
and add
RT on ice
Pre-incubate at
42°C or 50°C
then add RT
Add
RT on ice
65°C
42°C
or
50°C
0°C
Add RT
PrimeScript RT
(RT at 42°C)
Company L RT III
(RT at 50°C)
321
3
3
2
2
1 321
1
Figure 4: cDNA synthesis suppression by addition of RT to
non-denatured RNA.
Figure 3: Accuracy of various RTs.
RT Bases
sequenced
Error
(bases)
Error rate (%)
PrimeScript RT 201,297 7 0.0035
Company L RT II 166,227 9 0.0054
Company L RT III 161,409 11 0.0068
Company A RT 132,962 13 0.0098
M-MLV (Takara) 144,504 15 0.0104
Control (PCR only)
156,188 1 0.0006
Figure 2: First strand cDNA synthesis with various RTs.
kb
12 -
10 -
8.4 -
6.4 -
4.4 -
2.0 -
1.0 -
1 2 3 4 5 6 7 8
Figure 2:
First strand cDNA synthesis with
various RTs
1: PrimeScript RT 200 U
2: PrimeScript RT 10 0 U
3: Company L RT III
4: Company P RT
5: Company A RT
6: Company T RT
7: Company R RT
8: Company Q RT
RT-q PCR REAGENTS
3
To further assess the robustness of these enzymes, the
ability of each RT to synthesize cDNA after prolonged
incubation on ice was analyzed. RT was incubated with
denatured RNA template on ice for 0, 5, or 15 minutes
before proceeding with reverse transcription (Figure
5). PrimeScript RT produced high yield of dystrophin
cDNA even after a 15 minutes incubation on ice, whereas
synthesis of cDNA by Company L’s RT was completely
suppressed under the same condtions.
These experiments demonstrate the ability of PrimeScript
RT to synthesize cDNA under a variety of protocol
conditions. Taken together, the data indicate that
PrimeScript RT provides greater flexibility in workflow,
which can be particularly advantageous for medium- and
high-throughput studies.
Experimental Examples
Example 1: Reverse transcription
of template RNA that has strong
secondary structure
[Procedure] A 418 bp cDNA product
from 28S ribosomal RNA, which
contains strong higher order structure
(Figure 6, left panel), was synthesized
using either PrimeScript RT or Company
L’s RT. Total RNA from HL60 cells
was used as template, and a gene-
specific primer was used for cDNA
synthesis. PrimeScript RT reactions
were performed at 42°C and Company
L reactions were performed at 50°C
or 55°C using each manufacturer’s
recommended protocols. The efficiency
of cDNA synthesis was evaluated using PCR.
[Result] As shown in Figure 6, PrimeScript RT had the
highest sensitivity and transcription efficiency. These results
are consistent with the high RT efficiency of PrimeScript RT
for RNA that contains strong secondary structure.
Example 2: Reverse Transcription of GC-rich RNA
[Procedure] cDNA synthesis from human heart total
RNA primed with oligo-dT was performed using either
PrimeScript RT or Company L’s RT. The reactions were
assembled and performed according to each manufacturer’s
recommended conditions. A 520 bp fragment of the APOE
gene (~75% GC content) was amplified by PCR.
[Result] A higher yield of APOE cDNA was obtained using
PrimeScript RT (Figure 7). In contrast, Company L’s RT gave
lower yield of cDNA from the GC-rich APOE template.
Reverse transcription of RNA with high secondary structure
Marker: 100 bp DNA Ladder
PCR on cDNA template synthesized from 500 fg; 5 pg; 500pg; 5 ng; 50 ng; 500 ng
of total RNA (from left to right).
418
bp
PrimeScript RT
(42°C)
RT III (50°C)
Company LCompany L
RT III (55°C)
Specific
RT Primer
F
:
Forward Primer
R
:
Reverse Primer
R
F
Figure 7: RT-PCR of GC-rich region (APOE gene)
M 1 2 3 4 5 M 1 2 3 4 5 M
PrimeScript RT Company L RT III
Template cDNA amount used (corresponding to total RNA)
1 : 0 ng
2 : 0.125 ng
3 : 1.25 ng
4 : 12.5 ng
5 : 125 ng
M : 10 0 bp ladder
Figure 5: cDNA synthesis after extended pre-incubation on ice.
Figure 6: Reverse transcription of RNA with high secondary structure.
Figure 7: RT-PCR of GC rich region (APOE gene).
Figure 5: cDNA synthesis after extended pre-incubation on ice
RT left on ice for:
RNA denaturation:
PrimeScript RT
+ + + -
0 5 15 0
+ + + -
0 5 15 0
min.
Company L RT III

RT-q PCR REAGENTS
4
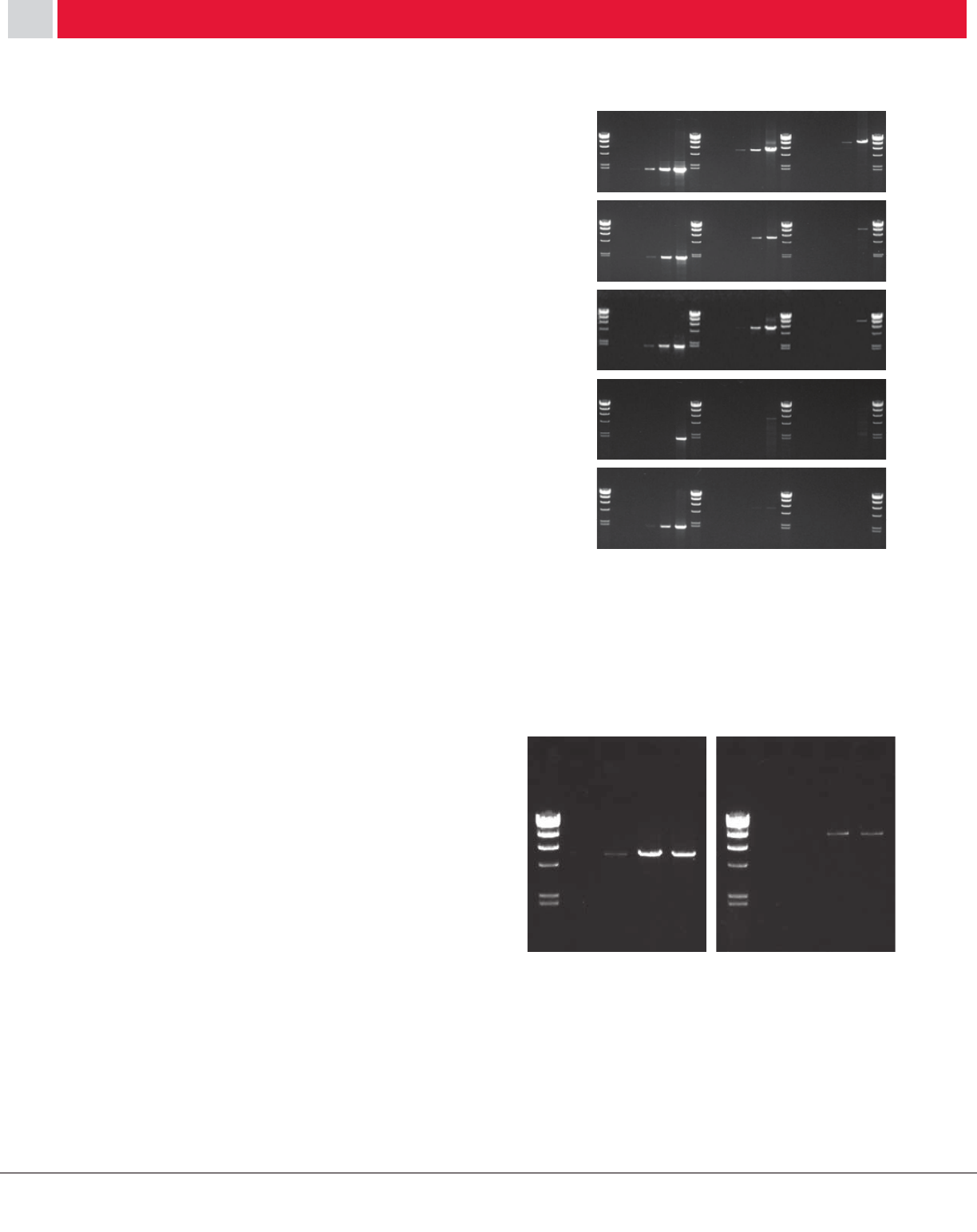
Example 3: Long RT-PCR
[Procedure] cDNA was synthesized from total RNA from
HL60 cells with oligo-dT primers using either PrimeScript
RT or one of four other commercially-available RTs. The
reactions were assembled and performed according to each
manufacturer’s recommended conditions. Dystrophin cDNA
products (2 kb, 6 kb, or 12kb) were amplified by PCR.
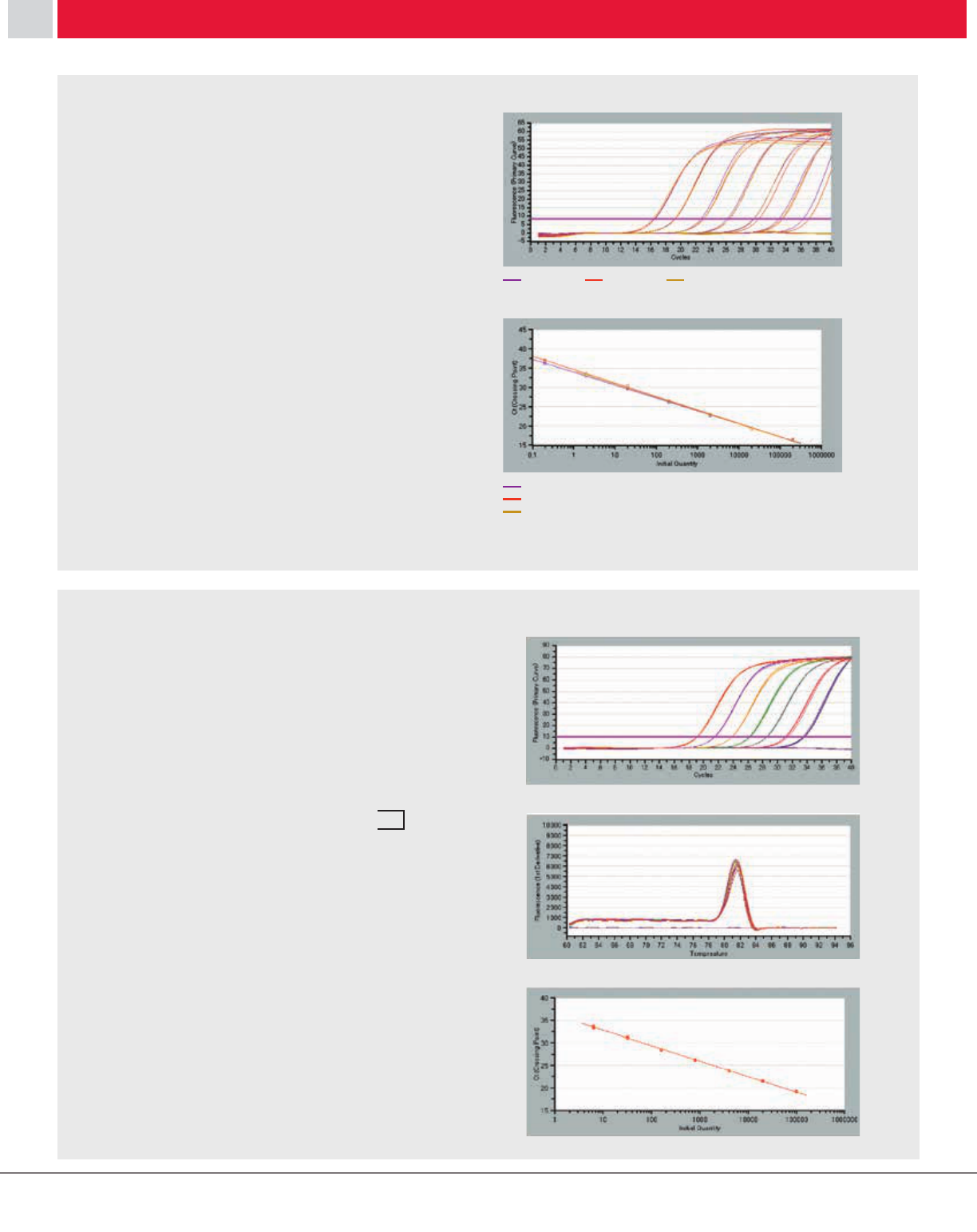
[Result] As shown in Figure 8, a higher yield was generally
observed for the PrimeScript RT-generated cDNA samples,
in particular the 6 kb and 12 kb cDNA products which could
not be amplified from cDNA synthesized with most of
the other commercially available enzymes. These results
indicate that PrimeScript RT provides superior yield of long
cDNA products.
Example 4: RT-PCR using random primers
[Procedure] cDNA was synthesized from 0 ng, 50 ng, 500
ng, or 1 µg of input RNA (human heart total RNA, Clontech
Cat. # 636532) using PrimeScript RT with random primers.
After cDNA synthesis, 6 kb or 12 kb fragments of the Dys-
trophin gene were amplified by PCR.
[Result] As shown in Figure 9, PrimeScript RT was capable
of synthesizing long cDNA (6 kb and 12 kb) with random
primers.
Description of Select PrimeScript Kits
PrimeScript 1st strand cDNA Synthesis Kit
This kit (Clontech Cat. # 6110A) contains all components
required for first-strand cDNA synthesis. It is designed for
long and sensitive first strand cDNA synthesis from total
RNA or polyA
+
RNA using PrimeScript RT. cDNA prepared
using this kit can be used for second-strand cDNA synthe-
sis, PCR amplification, or other applications such as cDNA
library preparation.
PrimeScript RT-PCR Kit
Perform two-step RT-PCR efficiently on any RNA template
with this kit (Clontech Cat. # RR014A), which includes an op-
timized combination of
TaKaRa Ex Taq
®
HS and PrimeScript
RT. The combination of enzymes allows excellent extension
and highly efficient amplification. This kit provides all of the
reagents required for reverse transcription and PCR (Figure
10), and is best suited for general RT-PCR applications.
PrimeScript One-Step RT-PCR Kit
The PrimeScript One Step RT-PCR Kit (Clontech Cat. # RR055A)
combines PrimeScript RT and
TaKaRa Ex Taq
HS for excel-
lent performance and ease of use. This kit allows all re-
agents for reverse transcription and PCR to be assembled
in one tube, making handling simple and greatly minimiz-
ing the risk of contamination (Figure 11).
Figure 9: Reverse transcription using random 6-mer primers
RT reaction: 50 min.
1: 0
2: 50 ng
3: 500 ng
4: 1 µg
M: λ-Hind III
Dystrophin 6 kb
M 1 2 3 4
Dystrophin 12 kb
M 1 2 3 4
Figure 8: Comparison of RT-PCR using various RTases
PrimeScript
RT
Company L
RT III
Company L
RT II
Company T
RT
Company Q
RT
* PCR was performed under the same conditions (30 cycles)
Template cDNA amount used in PCR (corresponding to total RNA)
1: 0; 2: 125 pg; 3: 1.25 ng; 4: 12.5 ng; 5: 125 ng; M: l-Hind III
Dys 2 kb Dys 6 kb Dys 12 kb
M 12345M12 345M1 2345M
Figure 8: Comparison of RT-PCR using various RTs.
Figure 9: Reverse transcription using random 6-mer primers.

RT-q PCR REAGENTS
5
PrimeScript High Fidelity RT-PCR Kit
This two-step RT-PCR kit (Clontech Cat. # R022A) provides
outstanding accuracy by combining PrimeSTAR Max
DNA polymerase and PrimeScript RT. PrimeSTAR Max is
a hot-start PCR enzyme with the highest accuracy of any
commercially available enzyme. The PrimeScript High
Fidelity RT-PCR Kit is compatible with a broad range of
template RNA levels. This kit is recommended for any
RT-PCR application that requires high accuracy.
PrimeScript RT Reagent Kit (Perfect Real Time)
for qRT-PCR
This kit (Clontech Cat. # RR037A) is designed for two
step real time RT-PCR. It allows easy, rapid preparation of
template cDNA using PrimeScript RT, followed by efficient
cDNA amplification using SYBR
Premix Ex Taq
™ (Cat.
#RR420A) or
Premix Ex Taq
(Clontech Cat. #RR390A)
(Figure 12).
PROTOCOL
20 µl RT reactions were performed using
various amounts of total RNA. Then, 2 µl of
each RT reaction was used in a 50-µl PCR
reaction.
RESULT
The PrimeScript RT-PCR Kit performed
better than the previous generation Takara
RT-PCR Kit with AMV RT. This result is likely
due to the higher RT efficiency imparted by
PrimeScript RT versus conventional AMV
RT
.
Figure 10. Comparison between PrimeScript RT-PCR Kit and a kit containing a conventional AMV RT.
Figure 11. Performance of PrimeScript One Step RT-PCR Kit and One Step RT-PCR kits from other suppliers for products up to 8kb long.
PROTOCOL
Various sizes of targets were amplified by RT-PCR. 1 µg of total human
RNA was used as a template for cDNA synthesis. Each RT-PCR reaction
was performed according to the supplier’s recommended conditions.
RESULT
The PrimeScript One Step RT-PCR Kit resulted in better amplification of
longer products (up to 8 kb) than the other kits tested.
PrimeScript
One Step RT-PCR Kit
M123456M
Company L
RT-PCR system III
M123456M
Company R
RT-PCR Kit
M123456M
Company Q
RT-PCR Kit
M123456M
Template : human total RNA
Lane 1 : TFR 0.5 kb
2 : CCND2 2.1 kb
3 : CCND2 2.8 kb
4 : TFR 4.4 kb
5 : Dystrophin 6 kb
6 : Dystrophin 8 kb
M : l-Hind III digest
PrimeScript RT-PCR Kit
2.2 kb 4.4 kb 6 kb 12 kb
One-Step RNA PCR Kit (AMV)
PrimeScript RT-PCR Kit
One-Step RNA PCR Kit (AMV)
Target: TFR
Template: total RNA from HL60 cells
0; 1; 10; 100; 10 00 ng (left to right)
Target: Dystrophin
Template: total RNA from human heart
0; 1; 10; 100; 10 00 ng (left to right)
2.2 kb 4.4 kb 6 kb 12 kb
RT-q PCR REAGENTS
6
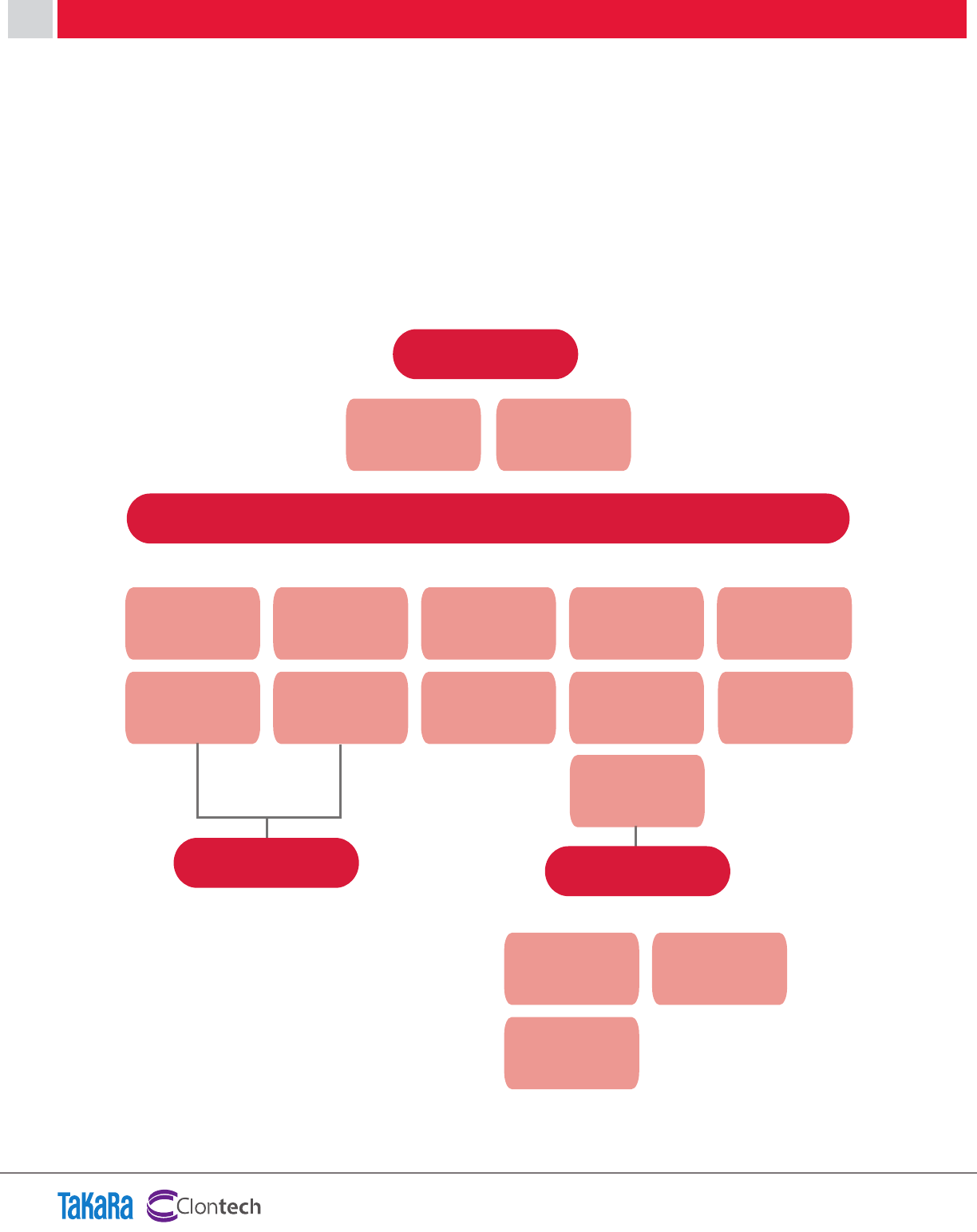
Figure 12. Performance of PrimeScript RT reagent Kit (Perfect Real Time) with various RT reaction lengths.
Figure 13. Performance of Real-Time One Step
RT-PCR with SYBR Green I detection.
PROTOCOL
[RT]
Template: mouse liver total RNA 2 pg —2 µg
or distilled water (negative control)
Volume: 10 µl
Primer: Random 6mers
Reaction conditions: 15, 30, or 60 minutes at 37°C;
5 seconds at 85°C; 4°C.
[Real-Time PCR]
Reagent used:
SYBR Premix Ex Taq
(Perfect Real Time)
Template: 2 µl of above the RT reaction
Final volume: 25 µl
Target gene: mouse Actb
Instrument: Thermal Cycler Dice™ Real Time System
(not available in all geographic regions)
RESULT
The results obtained using various RT reaction times
indicate excellent efficiency over a wide range of
template concentrations.
Amplification curve
Standard curve
RT 15 min. Rsq: 0.999 Eff: 98.7% Y = 3.552×LOG (X) + 33.94
RT 30 min. Rsq: 0.999 Eff: 93.3% Y = 3.495
×LOG (X) + 34.61
RT 60 min. Rsq: 0.999 Eff: 95.2% Y = 3.441
×LOG (X) + 34.28
RT 15 min. RT 30 min. RT 60 min.
PROTOCOL
Reagent: One Step SYBR PrimeScript RT-PCR Kit
(Perfect Real Time)
Template: Mouse liver total RNA 6.4 pg—100 ng
Target gene: rat Rplp2 (ribosomal protein, large P2)
Instrument: Thermal Cycler Dice Real Time System
(not available in all geographic locations)
Reaction conditions:
1) RT step 42°C 5 min.
95°C 10 sec.
2) PCR step 95°C 5 sec.
40 cycles
60°C 30 sec.
3) Melting step 60°C to 94°C
RESULT
Rplp2 was detected with total RNA inputs ranging from
6.4 pg
—100 ng. Good linearity of the standard curve
was obtained. These results indicate that the One Step
SYBR PrimeScript RT-PCR Kit (Perfect Real Time) provided
accurate quantification across the range of input RNA
amounts tested.
RSq
=
1.000
Eff
=
95.0%
Amplification curve
Denaturation curve
Standard curve
RT-q PCR REAGENTS
7
One Step SYBR PrimeScript RT-PCR Kit II (Perfect
Real Time) & One Step PrimeScript RT-PCR Kit
(Perfect Real Time)
These systems are designed for one-step RT-PCR using
PrimeScript RT for cDNA synthesis and
TaKaRa Ex Taq
HS
for PCR amplification.
Two kits are available: one for detection with SYBR Green I
(Clontech Cat. #RR086A), and the other for detection with
a TaqMan
®
probe (Clontech Cat. #RR064A). Because these
systems provide excellent amplification rate and reaction
specificity, they are highly recommended when working
with small amounts of RNA or RNA viruses (Figure 13).
Choosing the Best PrimeScript Kit for Your Experiment
qPCR
SYBR Premix Ex Taq
DNA Polymerase
(Tli RNase H+)
(RR420)
Premix Ex Taq
(Probe qPCR)
(RR390)
SYBR Premix Ex Taq
DNA Polymerase II
(Tli RNase H+)
(RR820)
SYBR Probe
PrimeScript RT
Reagent Kit
(Perfect Real Time)
(RR037)
Two-Step RT-qPCR
Reverse Tr anscription
PrimeScript RT
Master Mix
(RR036)
One Step PrimeScript
RT-PCR Kit
(Perfect Real Time)
(RR064)
One-Step RT-qPCR
PrimeScript RT
Reagent Kit gDNA Eraser
(Perfect Real Time)
(RR047)
PrimeScript
RT-PCR Kit
(RR014)
Two-Step RT-PCR
PrimeScript
One Step RT-PCR Kit,
Ver. 2
(RR055)
One-Step RT-PCR
PrimeScript
One Step RT-PCR Kit,
Ver. 2 (Dye Plus)
(RR057)
PrimeScript
High Fidelity RT-PCR
Kit (R022)
CellAmp™ Direct RNA
Prep Kit for RT-PCR
(Real Time)
(3732)
PrimeScript
Reverse Transcriptase
(2680)
PrimeScript
1st cDNA Synthesis
Kit (6110)
End-Point PCR
cDNA Synthesis
One Step SYBR
PrimeScript II RT-PCR Kit
(Perfect Real Time)
(RR086)
CellAmp
Whole
Transcriptome
Amplification Kit (Real
Time), Ver. 2 (3734)
Sample Preparation
(for Real-Time RT-PCR)

RT-q PCR REAGENTS
8
PRODUCTS
Cat. # Product Package Size
2680A
2680B
PrimeScript Reverse Transcriptase 10,000 Units
40,000 Units
6110A
6110B
PrimeScript 1st strand cDNA Synthesis Kit 50 rxns
200 rxns
R022A
R022B
PrimeScript High Fidelity RT-PCR Kit 50 rxns
200 rxns
RR014A
RR014B
PrimeScript RT-PCR Kit 50 rxns
200 rxns
RR036A
RR036B
PrimeScript RT Master Mix (Perfect Real Time) 200 rxns
800 rxns
RR037A
RR037B
PrimeScript RT Reagent Kit (Perfect Real Time) 200 rxns
800 rxns
RR047A
RR047B
PrimeScript RT Reagent Kit with gDNA Eraser
(Perfect Real Time)
200 rxns
800 rxns
RR055A
RR055B
PrimeScript One Step RT-PCR Kit, Ver. 2 50 rxns
200 rxns
RR057A
RR057B
PrimeScript One Step RT-PCR Kit, Ver. 2 (Dye Plus) 50 rxns
200 rxns
RR064A
RR064B
One Step PrimeScript RT-PCR Kit (Perfect Real Time) 100 rxns
500 rxns
RR086A
RR086B
One Step PrimeScript RT-PCR Kit II (Perfect Real Time) 100 rxns
500 rxns
3734 CellAmp Whole Transcriptome Amplification Kit
(Real Time), Ver. 2
100 rxns
RELATED PRODUCTS
Cat. # Product Package Size
R050A PrimeSTAR GXL DNA Polymerase 250 Units
R045A PrimeSTAR Max DNA Polymerase 100 rxns
RR390A Premix Ex Taq (Probe qPCR) 200 rxns
RR420A SYBR Premix Ex Taq (Tli RNase H Plus) 200 rxns
RR820A SYBR Premix Ex Taq II (Tli RNase H Plus) 200 rxns
Please check www.clontech.com for current pricing.
Notice to Purchaser
Your use of these products and technologies is subject to compliance with any applicable licensing requirements described on the product’s web page at http://www.clontech.com.
It is your responsibility to review, understand and adhere to any restrictions imposed by such statements.
Clontech products are to be used for research purposes only. They may not be used for any other purpose, including, but not limited to, use in drugs, in vitro diagnostic purposes,
therapeutics, or in humans. Clontech products may not be transferred to third parties, resold, modified for resale, or used to manufacture commercial products or to provide a service
to third parties without prior written approval of Clontech Laboratories, Inc.
